The Cytoplasm and Its Organelles
The cytoplasm surrounds the
nucleus, and it is in the cytoplasm that the work of the cell takes place.
Cytoplasm is essentially a colloidal solution that contains water,
electrolytes, suspended proteins, neutral fats, and glycogen molecules.
Although not contributing to the cell’s function, pigments may also accumulate
in the cytoplasm. Some pigments, such as melanin, which gives skin its color,
are normal constituents of the cell. Bilirubin is a normal major pigment of
bile; its excess accumulation in cells is evidenced clinically by a yellowish discoloration
of the skin and sclera, a condition called jaundice.
Embedded in the cytoplasm are
various organelles, which function as the organs of the cell. These
organelles include the ribosomes, ER, Golgi complex, mitochondria, and
lysosomes.
Ribosomes
The ribosomes serve as sites of
protein synthesis in the cell. They are small particles of nucleoproteins (rRNA
and proteins) that are held together by a strand of mRNA to form polyribosomes
(also called polysomes). Polyribosomes exist as isolated clusters of
free ribosomes within the cytoplasm (Fig.
4.2) or attached to the membrane of the ER. Whereas free ribosomes are involved in the synthesis of
proteins, mainly enzymes that aid in the control of cell function, those
attached to the ER translate mRNAs that code for proteins secreted from the
cell or stored within the cell (e.g., granules in white blood cells).
Endoplasmic Reticulum
The ER is an extensive system of
paired membranes and flat vesicles that connect various parts of the inner cell
(see Fig. 4.2). Between the paired ER membranes is a fluid-filled space called
the matrix. The matrix connects the space between the two membranes of the
nuclear envelope, the cell membrane, and various cytoplasmic organelles. It
functions as a tubular communication system for transporting various substances
from one part of the cell to another. A large surface area and multiple enzyme
systems attached to the ER membranes also provide the machinery for a major
share of the cell’s metabolic functions.
Two forms of ER exist in
cells—rough and smooth. Rough ER is studded with ribosomes attached to
specific binding sites on the membrane. Proteins produced by the rough ER are
usually destined to become components of lysosomes or other organelles,
incorporated into cell membranes, or leave the cell as a secretory protein. The
rough ER segregates these proteins
from other components of the cytoplasm and modifies their structure for a specific function. For example, the
synthesis of both digestive enzymes by pancreatic acinar cells and plasma
proteins by liver cells takes place in the rough ER. All cells require a rough
ER for the synthesis of lysosomal enzymes.
The smooth ER is free of
ribosomes and is continuous with the rough ER. It does not participate in
protein synthesis; instead, its enzymes are involved in the synthesis of lipid molecules,
regulation of intracellular calcium, and metabolism and detoxification of
certain hormones and drugs. It is the site of lipid, lipoprotein, and steroid
hormone synthesis. The sarcoplasmic reticulum of skeletal and cardiac muscle
cells is a form of smooth ER. Calcium ions needed for muscle contraction are
stored and released from cisternae of the sarcoplasmic reticulum. The smooth ER
of the liver is involved in glycogen storage and metabolism of lipid-soluble
drugs.
The processing ability of the ER is
not unlimited. If proteins accumulate in the ER faster than they can be processed,
the cell is said to experience “ER stress,” and signaling mechanisms kick in to
slow protein production and restore homeostasis. If these homeostatic responses
fail, cell death (apoptosis) can result. Defects in the response to ER stress
can cause inflammation and even cell death. They have been implicated in
inflammatory bowel disease, a genetic form of diabetes mellitus, and a disorder
of skeletal muscle known as myositis, as well as many other diseases.
Golgi Complex
The Golgi apparatus, sometimes
called the Golgi complex, consists of four or more stacks of thin,
flattened vesicles or sacs (see Fig. 4.3). These Golgi bodies are found near
the nucleus and function in association with the ER. Substances produced in the
ER are carried to the Golgi complex in small, membrane-covered transfer
vesicles. Many cells synthesize proteins that are larger than the active
product. The Golgi complex modifies these substances and packages them into
secretory granules or vesicles. Insulin, for example, is synthesized as a
large, inactive proinsulin molecule that is cut apart to produce a smaller,
active insulin molecule within the Golgi complex of the beta cells in the
pancreas. In addition to producing secretory granules, the Golgi complex is
thought to produce large carbohydrate molecules that combine with proteins
produced in the rough ER to form glycoproteins. Recent data suggest that the
Golgi apparatus has yet another function: it can receive proteins and other
substances from the cell surface by a retrograde transport mechanism. Several bacterial
toxins, such as Shiga and cholera toxins, and plant toxins, such as ricin, that
have cytoplasmic targets have exploited this retrograde pathway.
Lysosomes and Peroxisomes
Lysosomes can be viewed as the digestive
system of the cell. These small, membrane-enclosed sacs
contain powerful hydrolytic enzymes. These enzymes can break down excess and
worn-out cell parts as well as foreign substances that are taken into the cell. All of the lysosomal
enzymes are acid hydrolases, which
means they require an acidic environment. The lysosomes provide this
environment by maintaining a pH of approximately 5 in their interior. The pH of
the cytoplasm, which is approximately 7.2, serves to protect other cellular
structures from this acidity. Primary lysosomes are membrane-bound
intracellular organelles that contain a variety of hydro- lytic enzymes that
have not yet entered the digestive process. They receive their enzymes as well
as their membranes from the Golgi apparatus. Primary lysosomes become secondary
lysosomes after they fuse with membrane-bound vacuoles that contain
material to be digested. Lysosomes break down phagocytosed material by either
heterophagy or autophagy (Fig. 4.4).
Heterophagy refers to digestion of
an exogenous substance phagocytosed from the cell’s external environment. An
infolding of the cell membrane takes external materials into the cell to form a
surrounding phagocytic vesicle, or phagosome. Primary lysosomes then
fuse with phagosomes to form secondary lysosomes. Heterophagocytosis is most
common in phagocytic white blood cells such as neutrophils and macrophages.
Autophagy involves the segregation and disposal of damaged cellular organelles,
such as mitochondria or ER, which the lysosomes must remove if the cell’s
normal function is to continue. Autophagocytosis is most pronounced in cells undergoing atrophy. Although enzymes in
the secondary lysosomes can break
down most proteins, carbohydrates, and
lipids to their basic constituents, some materials remain undigested. These
undigested materials may remain in the cytoplasm as residual bodies or
are extruded from the cell by exocytosis. In some long-lived cells, such as
neurons and heart muscle cells, large quantities of residual bodies accumulate
as lipofuscin granules or age pigment. Other indigestible pigments, such as
inhaled carbon particles and tattoo pigments, also accumulate and may persist
in residual bodies for decades.
Lysosomes play an important role in
the normal metabolism of certain substances in the body. In some inherited
diseases known as lysosomal storage diseases, a specific lysosomal
enzyme is absent or inactive, in which case the digestion of certain cellular
substances (e.g., glucocerebrosides, gangliosides, sphingomyelin) does
not occur.7 As a result, these substances accumulate in the cell. In Tay-Sachs
disease, an autosomal recessive disorder, hexosaminidase A, which is the
lysosomal enzyme needed for degrading the GM ganglioside found in nerve
cell membranes, is deficient. Although
GM ganglioside accumulates in many
tissues, such as the heart, liver,
and spleen, its accumulation in the nervous system and retina of the eye causes
the most damage.7 There are multiple lysosome storage diseases, and new
guidelines are being developed by the American College of Medical Genetics
regarding diagnostic criteria and management for Fabry, Gaucher, and
Niemann-Pick A/B disease; glycogen storage disease type II; globoid cell
leukodystrophy; metachromatic leukodystrophy; and mucopolysaccharidoses types.
Smaller than lysosomes, spherical
membrane-bound organelles called peroxisomes contain a special enzyme
that degrades peroxides (e.g., hydrogen peroxide). Unlike lysosomes,
peroxisomes are not formed by the Golgi apparatus. Peroxisomes are
self-replicating like mitochondria and are initially formed by proteins produced by free ribosomes.
Peroxisomes function in the control
of free radicals. Unless degraded,
these highly unstable chemical compounds would otherwise damage other
cytoplasmic molecules. For example, catalase degrades toxic hydrogen peroxide
molecules to water. Peroxisomes also contain the enzymes needed for breaking
down very–long-chain fatty acids, which mitochondrial enzymes ineffectively
degrade. In liver cells, peroxisomal
enzymes are involved in the formation of
the bile acids.
Proteasomes
Three major cellular mechanisms are
involved in the break- down of proteins, or proteolysis. One of these is
by the previously mentioned endosomal–lysosomal degradation. Another cytoplasmic
degradation mechanism is the caspase pathway that is involved in
apoptotic cell death. The third method of proteolysis occurs within an
organelle called the proteasome. Proteasomes are small organelles
composed of protein complexes that are thought to be present in both the
cytoplasm and the nucleus. This organelle recognizes misformed and misfolded
proteins that have been targeted for degradation, including transcription
factors and the cyclins that are important in controlling the cell cycle. It has
been suggested that as much as one third of the newly formed polypeptide chains
are selected for proteasome degradation because of quality-control mechanisms
in the cell.
Mitochondria
The mitochondria are literally the
“power plants” of the cell because they transform organic compounds into energy
that is easily accessible to the cell. They do not make energy but extract it
from organic compounds. Mitochondria contain the enzymes needed for capturing
most of the energy in foodstuffs and converting it into cellular energy. This
multistep process is often referred to as cellular respiration because
it requires oxygen. Cells store most of this energy as high-energy phosphate
bonds in compounds such as adenosine triphosphate (ATP), using it to power the
various cellular activities. Mitochondria are found close to the site of energy
consumption in the cell (e.g., near the myofibrils in muscle cells). The
number of mitochondria in a given cell type varies by the type of activity the
cell performs and the energy needed to undertake this activity. For example, a
dramatic increase in mitochondria occurs in skeletal muscle repeatedly
stimulated to contract.
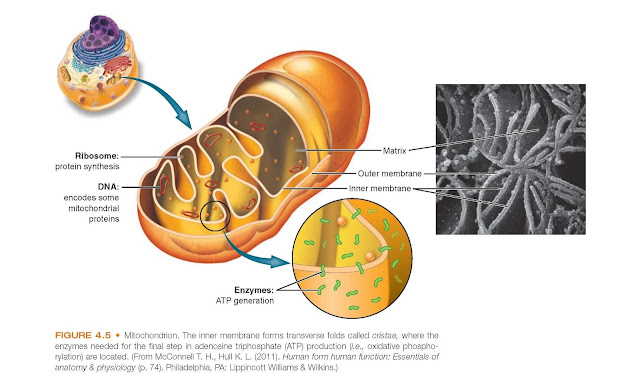
Mitochondria are composed of two
membranes: an outer membrane that encloses the periphery of the mitochondrion
and an inner membrane that forms shelflike projections, called cristae (Fig.
4.5). The narrow space between the outer and inner membranes is called the intermembrane
space, whereas the large space enclosed by the inner membrane is termed the
matrix space. The outer mitochondrial membrane contains a large number
of transmembrane porins, through which water-soluble molecules may pass.
Because this membrane is relatively permeable to small molecules,
including proteins, the contents
of the intermembrane space resemble that of the cytoplasm. The inner membrane
contains the respiratory chain enzymes and transport proteins needed for the
synthesis of ATP. In certain regions, the outer and inner membranes contact
each other, these contact points serve as pathways for proteins and small
molecules to enter and leave the matrix space.
Mitochondria contain their own DNA
and ribosomes and are self-replicating. Mitochondrial DNA (mtDNA) is found in
the mitochondrial matrix and is distinct from the chromosomal DNA found in the
nucleus. Also known as the “other human genome,” mtDNA is a double-stranded,
circular molecule that encodes the rRNA and tRNA required for
intramitochondrial synthesis of the proteins needed for the energy-generating
functions of the mitochondria. Although mtDNA directs the synthesis of 13 of
the proteins required for mitochondrial function, the DNA of the nucleus
encodes the structural proteins of the mitochondria and other proteins needed
to carry out cellular respiration.
mtDNA is inherited matrilineally (i.e.,
from the mother), thus
providing a basis for familial lineage studies. Mutations have been found in
each of the mitochondrial genes, and an understanding of the role of mtDNA in
certain diseases is beginning to emerge. Most tissues in the body depend to
some extent on oxidative metabolism and can therefore be affected by mtDNA
mutations.
Mitochondria also function as key
regulators of apoptosis or programmed cell death. The initiation of the
mitochondrial pathway for apoptosis results from an increase in mitochondrial
permeability and the subsequent release of proapoptotic molecules into the
cytoplasm. One of these proapoptotic molecules is cytochrome c, which is bound
by cardiolipin (a phospholipid). It is well known for its role in mitochondrial
respiration. In the cytosol, cytochrome c binds to a protein called apoptosis
activating factor-1, initiating the molecular events involved in the
apoptosis cascade. Other apoptotic proteins also enter the cytoplasm, where
they bind to and neutralize the various apoptotic inhibitors, whose normal
function is to block the apoptotic cascade. Both the formation of reac- tive
oxygen species (ROS) (e.g., peroxide) and the activation of the p53 tumor
suppressor gene by DNA damage or other means initiate apoptotic signaling
through the mitochondria. ROS has been determined to be the etiology of cell
injury to multiple diseases. Dysregulated apoptosis (too little or too much)
has been implicated in a wide range of diseases, including cancer, in which
there is an inappropriately low rate of apoptosis, and neurodegenerative
diseases, in which there is an
increased or excessive rate of apoptosis.





.webp)

_Page_049.jpg)