Invariant Natural
Killer T‐Cell Receptors Bridge Innate and Adaptive Immunity
The highly
variable nature of the TCR confers on the conventional T‐cell population the
ability to respond to an immense array of different antigens, with individual
T‐cells specific for a single antigen. Invariant natural killer T‐cells (iNKT)
are a unique subset of T‐cells that display a semi‐variant TCR that equips
individual iNKT cells with the ability to detect a broad array of microbial
lipid antigens, presented on CD1d antigen‐presenting molecules on
antigen‐presenting cells (APCs). Although conventional T‐cells are activated by
APCs that have first been activated by microbial antigen (in a process that
takes some time), iNKTs can respond directly to PAMPs, secreting cytokines and
presenting co‐stimulatory molecules in a manner more reminiscent of innate
immune cell PRR activation than T‐cell stimulation.
 |
Figure 4.15 Natural
killer T‐cells. (a) Schematic representation of type I and type II natural
killer T (NKT) cells. These two subsets use different variable (V) region gene
segments in the α and β chains of their T‐cell receptors (TCRs), and they
recognize different CD1d‐ restricted antigens. (b) The αβTCR is composed of two
chains, with the V domains containing the complementarity determining region
(CDR) loops. The CDR3 loops are encoded by multiple gene segments and also
contain nontemplated (N) regions, which add further diversity to the TCR
repertoire. The color coding is the same as that used for the type I NKT TCR in
(a). APC, antigen‐presenting cell; C, constant; D, diversity; J, joining.
(Reproduced with permis sion from the authors Rossjohn et al., (2012) Nature
Reviews Immunology 12, 845–857 © Nature Publishing Group.)
Although
conventional CD4+ T‐cells provide help to B‐ cells as part of an
adaptive immune response, iNKTs are unique in that they can provide help to
B‐cells in an innate and adaptive manner, with differing outcomes. iNKTs that
are activated by antigen presented on B‐cell C1d can directly license B‐cell
activation in a cognate, innate‐like manner, through co‐stimulation with CD40L
and the production of various cytokines, such as IFNγ and IL‐21. This leads to
a restricted form of B‐ cell activation, with plasmablast expansion, early
germinal center development, modest affinity maturation, and primary
class‐switched antibody production, but lacking the development of plasma cells
and B‐cell memory responses. Alternatively, iNKTs that have been activated by
DCs presenting antigen can drive full B‐cell activation in a noncognate, or
adaptive fashion, by enlisting the help of CD4+ T‐cells to license
B‐cells, driving the generation of mature germinal centers, robust affinity
maturation, the development of antibody‐producing plasma cells, and a B‐cell
memory response.
Antigen binding by iNKT receptors
Similar to
conventional T‐cells, the TCR of iNKTs consists of an α and β chain, with each
chain divided into a constant (C) and variable (V) region, with the variable
regions conferring ligand diversity. The variable domain of the TCR α chain is
split into V and J regions, whereas the β chain is encoded by V, D, and J
domains (Figure 4.15). Three complementarity determining regions (CDRs) are
situated within the V domains of both α and β chains and these regions enerate
the antigen‐binding site of the receptor (Figure 4.15).
There are
two main subsets of iNKT cells, type I and type II iNKT, with ligand binding by
type I TCRs exhibiting characteristics of innate PRRs, whereas type II iNKT
receptors share some similarity with conventional TCRs. Type I cells possess a
semi‐invariant TCR α chain (Vα24 Jα18), combined with a restricted β chain
which utilizes Vβ11. The CD3α loop plays a particularly important role for type
I NKT as deletion of this region in mice greatly impairs ligand binding. A
defining characteristic of type I iNKT over type II cells is their ability to
detect CD1d‐bound α‐galactosyl ceramide (αGalCer), a glycolipid originally
isolated from the marine sponge Agelas mauritianus and more recently
shown to be a component of Bacteroides bacteria which inhabit the human
gut. CD1d binding of αGalCer is typical of other α‐ linked glycolipids, with
the hydrophobic portion of the ligand buried in the two main binding regions of
CD1d, the A′ and F′ pockets, while the polar region is exposed to solvent.
Binding of
ligand by type I TCR is a relatively rigid affair, regardless of
the nature of
the ligand, and is
dominated by germline‐encoded regions in the semi‐variant α chain, with
assistance from β chain motifs. The type I iNKT TCR positions itself parallel,
above the F′ pocket of CD1d, in a manner similar to innate‐like PRRs, with the
β chain CDR2β binding a region above the F′ pocket. The αGalCer is bound
directly by the CDR1α loop and bridging of both CD1d and αGalCer by the CDR3α
loop stabilizes the inter action (Figure 4.16). The importance of the CDR2β
and CDR3α loops to ligand binding is illustrated by severely reduced ligand
binding in receptors bearing mutations to critical residues in these regions.
Many microbial ligands have been demonstrated for type I iNKT cells, including
α‐ glucosyldiacylglycerols from Streptococcus pneumoniae and
α‐galactosyldiacylglycerols from Borrelia burgdorferi. As with αGalCer,
the α their microbial origin as mammalian glycolipids have mainly β‐glycosidic
linkages.
Type II iNKT
cells do not respond to α‐glycolipids such as αGalCer, however, type II
receptors possess a more varied, oligoclonal repertoire than type I cells that
share features of both conventional and innate‐like T‐cells. Type II iNKT
specific for the self glycolipid antigen sulfatide have been found to play a
role in the regulation of various autoimmune disorders, includ ing
concanavalin A‐induced hepatitis (a mouse model of human autoimmune hepatitis)
and type 1 diabetes. The crystal structure of the type II Vα1Jα26–Vβ16Jβ2.1 TCR
bound to CD1d/sulfatide shows that, in contrast to the type I receptor, where
innate‐like germline‐encoded regions confer specificity, non‐germline‐encoded
regions, more similar to conventional TCRs, dominate type II iNKT ligand
binding (Figure 4.16). Although the type I receptor is orientated above the
CD1d F′ pocket, the type II TCR is positioned above the A′ pocket with both the
α and β chains making contact with CD1d in a diagonal orientation, similar to
MHC–TCR interactions. Additionally, the CDR3β loop of the type II receptor
determines specificity for sulfatide while the α chain of the type I TCR
interacts with ligand. The more diverse nature of type II TCR binding is
thought to confer on type II iNKT cells the ability to respond to a more varied
range of antigens, including sulfatides and other glycolipids, phospholipids,
and nonlipid antigens. The ability of type I and II iNKT TCRs to bind
CD1d‐presented antigen in mechanistically distinct ways illustrates the
impressive nature of mammalian immune systems in aried range of pathogen
products.
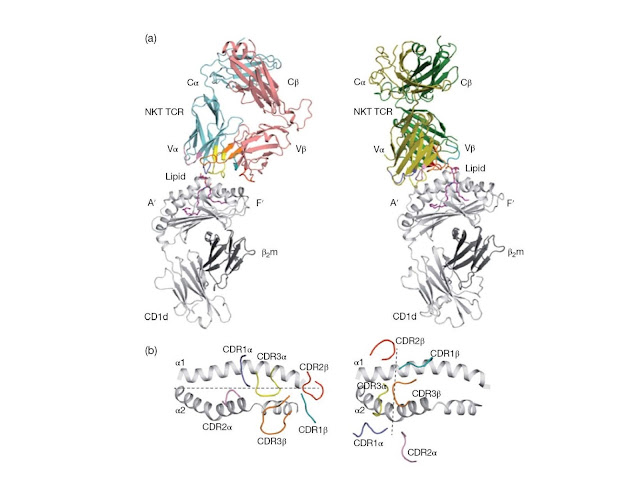 |
Figure 4.16 Structural comparison between
type I and type II NKT TCR–lipid–CD1d complexes. (a) The figure shows the
docking mode of the T‐cell receptor (TCR) in a type I natural killer T (NKT)
cell TCR–lipid–CD1d complex (left) and a type II NKT TCR–lipid–CD1d complex
(right). The CD1d antigen‐binding pockets are labelled A′ and F′. (b) The
figure shows the view looking down into the antigen‐binding groove of the two
complexes showing the parallel docking mode in the type I NKT–lipid–CD1d
complex (left), and the orthogonal docking mode in the type II NKT–lipid–CD1d
complex (right). Dashed lines represent the docking mode. β2m, β2‐microglobulin.
(Reproduced with permission from the authors Rossjohn et al., (2012) Nature
Reviews Immunology 12, 845–857 © Nature Publishing Group.)


.webp)

